

TOP

• The Role of ASO Institutions
Traditional antisense oligonucleotide (ASO)-based gene regulation (including silencing or enhancement of gene expression) targets mRNAs and can occur in either the nucleus or the cytoplasm. In the nucleus (targeting pre-mRNA precursors), regulation typically acts by altering polyadenylation, altering splicing events, or cleaving inter-nucleotide bonds, all of which occur during mRNA maturation (Figure 1). In the cytoplasm (targeting mature mRNAs), regulation usually acts through translational modulation or recruitment of RNaseH cleavage (Figure 2)
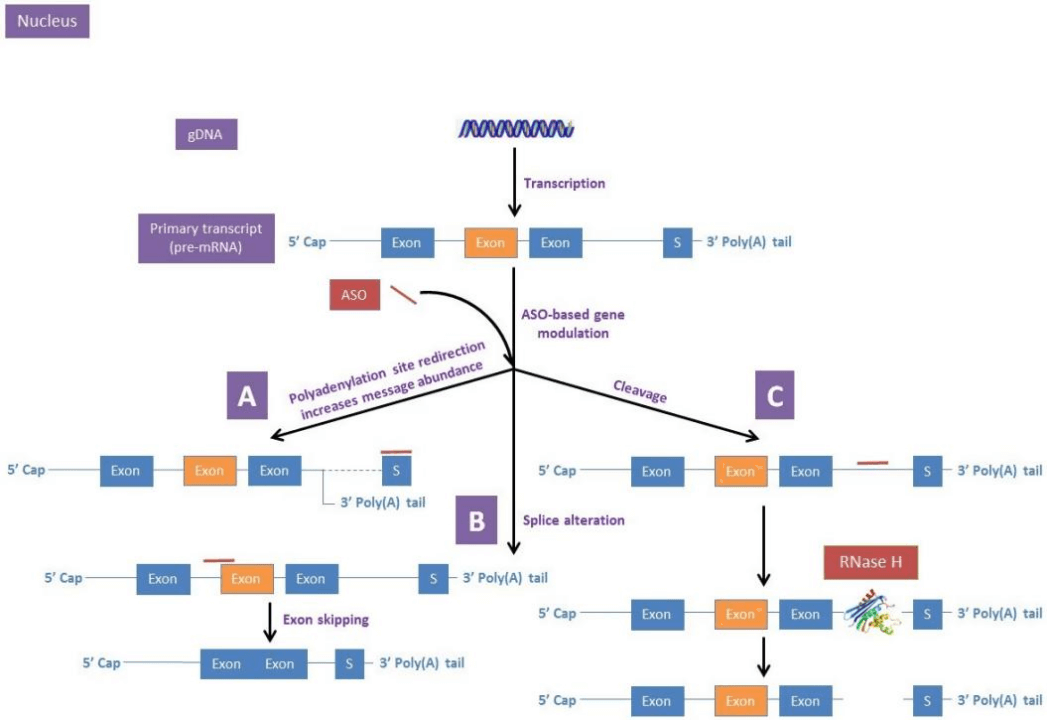
图1. Antisense oligonucleotide-based gene regulatory mechanisms in the nucleus of cells
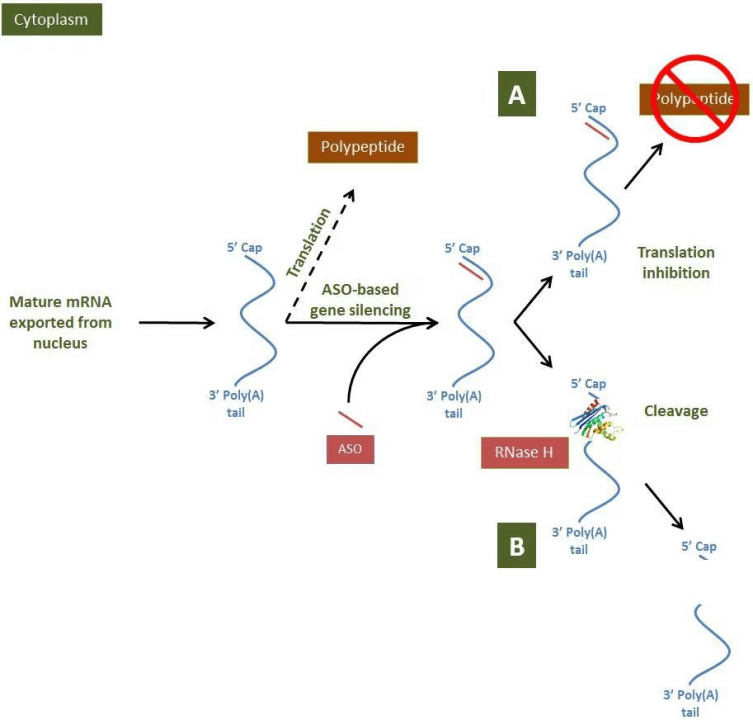
图2. Mechanisms of antisense oligonucleotide-based gene regulation in the cytoplasm of cells
Antisense oligonucleotides are recognized and hybridized to the target mRNA by base pairing. Resulting in the target mRNA being RNase H either in the nucleus or cytoplasm. Cleaved antisense oligonucleotides have been extensively studied for research and therapeutic purposes, and are therefore relatively well-studied in terms of regulatory mechanisms. rNase H uses magnesium ions as a cofactor to cleave mRNA strands in mRNA:DNA heterodimeric duplexes by hydrolyzing inter-nucleotide (phosphodiester) bonds.
• ASO Design Considerations
In principle, gene silencing should look like the selection of a sequence in the target mRNA, the procurement of complementary by base-pairing antisense oligonucleotides, their introduction into the system under study (either in vitro or in vivo), and then subsequent molecular biology to detect the effect of ASO action. However, designing antisense oligonucleotides requires consideration of many factors.
Hybridization site
According to base pairing rules, antisense oligonucleotides should hybridize to any region of the target mRNA sequence. However, mRNA folds into secondary or even tertiary structures, which may prevent antisense oligonucleotides from hybridizing. Therefore, the unfolded region of the mRNA should be selected as the hybridization site. There are wet-lab methods, such as RNase H localization, that can be used to predict accessible sites, and also to make preliminary judgments by predicting RNA secondary structure.
The frequency of unmethylated CpG dinucleotides is much higher in bacterial DNA than in vertebrate DNA. This is mainly due to the fact that CpG dinucleotides are underrepresented in vertebrate genomes and 80% of them are marked by methylation. Since CpG motifs in bacteria trigger the activation of B cells, natural killer cells, monocytes, and cytokines, whereas vertebrate CpG motifs do not, this may be one of the ways in which the immune system recognizes bacterial infection. Antisense oligonucleotides containing unmethylated CpG motifs stimulate the immune system in a manner similar to that of bacterial DNA and may be responsible for some of the effects reported in earlier antisense studies.
To avoid immune stimulation, if possible, design antisense oligonucleotides that do not contain CpG motifs, or at least the following extended motifs that generate the strongest immune responses.
The optimal length is usually 12 to 28 bases. Sequences shorter than 12 bases increase the probability of off-target hybridization, while sequences longer than 25 bases result in reduced cellular uptake.
The secondary structure of antisense oligonucleotides and the formation of oligonucleotide dimers should be examined, as both may interfere with hybridization to the target site sequence. If possible, design antisense oligonucleotides so that they have the weakest possible secondary structure and do not form dimers.
As mentioned above, it is critical to identify unfolded sites in the target mRNA and to ensure that antisense oligonucleotides do not have self-complementarity. However, these considerations alone are not sufficient to ensure proper hybridization. Various factors, such as phosphorothioates, can reduce the binding affinity of antisense oligonucleotides for the target site, which in turn can make attenuated knockdown inefficient. Third-generation antisense oligonucleotide modifications have been found not only to have nuclease-resistant properties but also to increase binding affinity.
The final unmodified antisense oligonucleotide sequence should be subjected to a BLAST search to ensure that any off-target hybridization (preferably none) does not interfere with antisense activity or cause unacceptable toxicity.
• Quality considerations
In principle, gene silencing should look like the selection of a sequence in the target mRNA, the procurement of complementary by base-pairing antisense oligonucleotides, their introduction into the system under study (either in vitro or in vivo), and then subsequent molecular biology to detect the effect of ASO action. However, there are many factors to consider when designing antisense oligonucleotides. For in vivo animal experiments, we recommend that antisense oligonucleotides undergo in vivo level purification, including ion exchange, endotoxin detection (to ensure that pyrogens are below the upper acceptable limit), and filtration (to reduce the number of contaminating colony-forming units to below the upper acceptable limit).